Everything we know about life and how it works ultimately comes down to the genetic blueprint, DNA. DNA, as organized in genes, are the basic building blocks of genomes and therefore of living cells. Genes are arguably one of the most important forms of currency we have to create life because each is the result of millions of years of selection and refinement to do its job. So, given the choice would organisms shed genes if they had the chance to keep them whole, or at least the pieces of them that carry out key functions, known as protein domains? In a recent collaborative paper with colleagues at the Université Pierre et Marie Curie in Paris, Genome Cooperative bioinformaticist Udi Zelzion, working with EOAS scientist Debashish Bhattacharya, helped answer this question.
It turns out that when the photosynthetic organelle, the plastid, originated in algae and plants through primary endosymbiosis over 1 billion years ago, the “host” inherited several thousand genes housed in the endosymbiont genome. Most of these genes however were discarded and others were moved to the nucleus of the algal host. At least that is how standard theory went. Using powerful network methods that can disentangle the reticulate (i.e., web-like) evolutionary history of genes, Méheust et al. (2016) discovered that dozens of endosymbiont genes were not lost at all, but rather recycled into new genes through domain fusions (so-called symbiogenetic, or S-genes). These protein similarity networks allow scientists to find genes that are made up of two unrelated parts, joined together to form a new chimeric sequence that does not exist anywhere else in the tree of life. Méheust et al. found 67 S-genes that are widely distributed among algae and plants and provide key functions related to tolerance to stress generated by photosynthesis, orienting cells to the light direction, and dealing with nutrient limitation. In fact some of the best studied genes in plants that, for example, play a role in repairing plastid genomic DNA after light stress, turn out to be S-genes.
So why did S-genes evolve? By studying their functions, Méheust et al. were able to surmise that endosymbiosis itself was such a stress to the algal host that it capitalized on the opportunity to recycle gene fragments to help it harness the new photosynthetic organelle. This work shows yet again how little we still understand about how genomes evolve on our planet. Genes are apparently too valuable to lose outright, if their component parts can be re-used to serve the host genome.
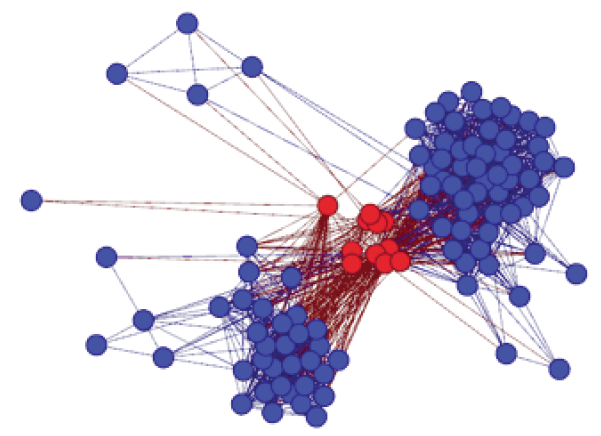
Above: A network diagram showing how two different protein domains (shown in blue circles) were united to give rise to a new S-gene (shown in red circles). This S-gene is found in Arabidopsis (AtHBP5) and in Chlamydomonas (SOUL3) where it functions in chlorophyll degradation in the plant and in coordinating response to light direction in the green alga.
Congratulations to Dr. Bhattacharya and his colleagues on this publication!
To learn more about this work, the full article may be found here.
—–
Méheust R, Zelzion E, Bhattacharya D, Lopez P, Bapteste E. 2016. Protein networks identify novel symbiogenetic genes resulting from plastid endosymbiosis. Proc Natl Acad Sci USA Early edition: www.pnas.org/cgi/doi/10.1073/pnas.1517551113.


